Newsletter / Research Activity Report / Hiroyoshi Matsumura
Newsletter Vol. 19 Drawing Worldwide Attention: Tackling Global Issues via Molecular Interface Control— An Interview with RARA Associate Fellow Hiroyoshi Matsumura on His Remarkable Consecutive Publications in Nature Communications
2025 / 10 / 21

2025 / 10 / 21
Achieving back-to-back publications in a top-tier journal and striving to open up the new field of “molecular interface control”
This edition of the newsletter features an interview with Professor Hiroyoshi Matsumura (RARA Associate Fellow; College of Life Sciences).

Prof. Matsumura has achieved outstanding research outcomes resulting in the presentation and publication of two papers in quick succession—one in 2023 and the other in 2025—in the internationally acclaimed journal Nature Communications (a peer-reviewed, open access journal published by Springer Nature).
Behind this accomplishment is a new topic that Prof. Matsumura is working on: biofunction science through the regulation of molecular interfaces. In conventional biochemistry, the dominant approach has been to enhance function by modifying the active sites of enzymes and protein, that is, the central regions where chemical reactions occur. However, Prof. Matsumura recognized the potential to enhance enzyme performance by manipulating molecular interfaces located away from active sites, and he is currently working on functional modification and the creation of entirely new functions.
He was the first in the world to elucidate the dynamic mechanisms of proteins involved in bacterial cell division, and he is also pursuing the development of completely new therapeutic drugs targeting drug-resistant bacteria such as MRSA, against which conventional antibiotics are ineffective.
Tackling global warming by improving the photosynthetic enzymes in plants
Another area currently drawing attention is the latest research findings of Prof. Matsumura, which suggest one possible solution to global warming. By enhancing the activity of RuBisCO (ribulose-1,5-bisphosphate carboxylase/oxygenase), the enzyme essential for carbon dioxide fixation in photosynthesis, he has found that the growth of the model plant Arabidopsis thaliana could be accelerated by approximately 1.3 times. This discovery holds promise for simultaneously addressing climate change mitigation through enhanced plant growth and global food security issues.
We spoke with Prof. Matsumura about the driving force behind his research, how he selects his research topics, his research philosophy, the breakthroughs he has achieved by collaborating with his students, and the keys to his success.
(The following is a summary of the conversation with Prof. Matsumura compiled by our writer.)
A discovery that unlocks the mystery of bacterial cell division
Below is a brief introduction to the two papers published in Nature Communications (for details, see Nat. Commun., 16, 5985 (2025) and Nat. Commun., 14, 4073 (2023)).
Bacterial infectious diseases continue to pose a serious global challenge. Although a wide range of antimicrobial drugs have been developed to combat these infections, the emergence of drug-resistant bacteria against which existing drugs are ineffective has created an urgent need for new types of antimicrobial drugs. Among these potential agents, a protein called FtsZ has attracted considerable attention as a candidate for new drug development, because it is present in many pathogenic bacteria and is essential for their proliferation.
FtsZ functions by assembling into elongated, filament-like structures resembling a string of beads, and many of its three-dimensional structures have been elucidated to date using X-ray diffraction. However, X-ray diffraction can only reveal the structure of a protein in a crystalline state. In a crystal, protein molecules are tightly packed together, and it has been pointed out that such structures may not accurately reflect the behavior of FtsZ as it moves dynamically inside living cells. For this reason, there has been a growing need to analyze its structure not in a crystal, but in solution, that is, an environment closer to that inside a cell.
To address this challenge, I initiated a joint research project with Professor Keiichi Namba, Assistant Professor Junzo Fujita, and Mr. Kazuki Kasai of the University of Osaka, and we decided to employ cryo-electron microscopy, a technique in which proteins in solution are rapidly frozen and observed using an electron beam. However, because FtsZ is a highly dynamic protein, determining its structure using cryo-electron microscopy has proven extremely difficult. Although researchers around the world have attempted this approach, it has repeatedly resulted in failure. In fact, we also had trouble at first and experienced a series of failures.
Therefore, we changed our approach and decided to try using artificial antibodies, or molecules that are artificially engineered to bind to specific target proteins. As part of our research into manipulating molecular interfaces located away from active sites, we all routinely worked on improving enzymes by attaching artificial antibodies to them. As a result, we were able to obtain artificial antibodies that bind to FtsZ relatively easily, and we successfully attached these antibodies to the molecular interface (i.e., the surface of the protein) of FtsZ. Doing this stabilized the previously volatile protein structure, as if a structural framework had been added, allowing us to observe in detail regions that had previously appeared only indistinctly (Fig. 1).
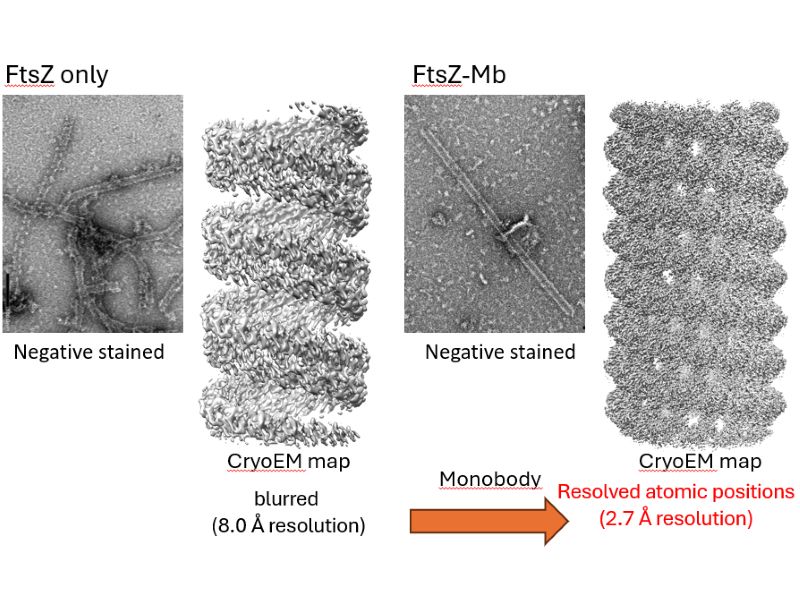
Fig. 1: The structure of FtsZ alone appeared only indistinctly (left), whereas the addition of the artificial antibody Mb enabled the structure to be visualized clearly down to the level of individual atomic positions (right).
It was astonishing to see something that had previously been invisible appear so clearly. When we first added the artificial antibody to FtsZ and observed it (the negatively stained FtsZ–Mb image in Fig. 1), we mistakenly thought we were seeing something other than FtsZ because it very much resembled the shape of a virus, and I even said, “This must be contamination.”
Anyhow, when we went ahead and conducted structural analysis, we found that binding this artificial antibody made it possible to analyze the bent structure of the FtsZ filaments. Building on insights gained from these experiments, we subsequently succeeded in analyzing the structure of straight FtsZ filaments as well (Fig. 2). When bacteria undergo cell division, FtsZ changes its structure from straight filaments to curved filaments (Fig. 3). In other words, because we were able to ascertain for the first time the structures of FtsZ filaments in both straight and curved states, we were able to elucidate how FtsZ filaments transition from a straight to a curved configuration during bacterial cell division.
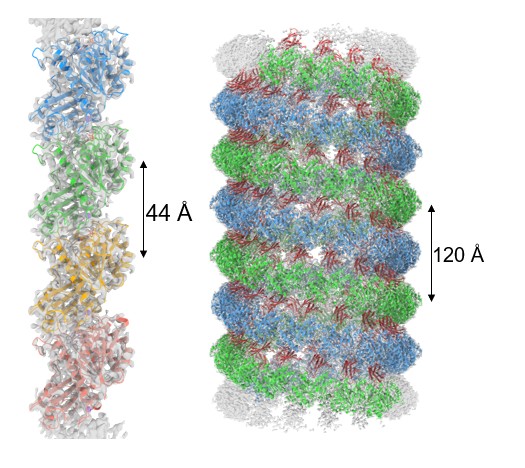
Fig. 2: Straight FtsZ filament (left) and curved (helical) FtsZ filament (right)
Elucidating the sophisticated mechanism that keeps filaments moving despite being densely packed
Cell division is a phenomenon in which the cell membrane is constricted inward (Fig. 3). To deform the membrane in this manner, the activity of FtsZ alone is not sufficient; other proteins must bind to the FtsZ filaments that serve as a scaffold, and these components must work in concert. The findings we reported in 2023 focused on observations of the FtsZ filaments alone, which means we could only understand part of the actual mechanism of cell division. To address this limitation, we conducted another study combining FtsZ with another protein, ZapA, which binds to FtsZ and aids its function.
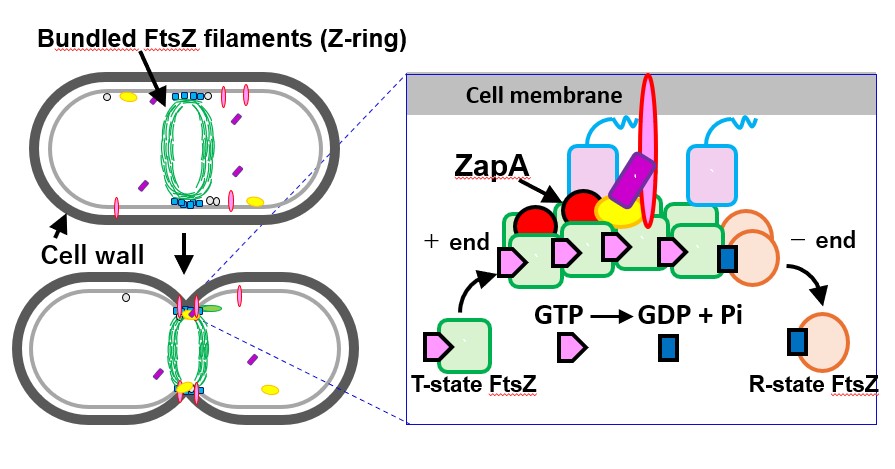
Fig. 3: Mechanism of cell division. Filamentous FtsZ assembled in a bead-like chain is thought to bind with other proteins such as ZapA, which cooperate with FtsZ to pull the cell membrane inward.
As described above, when bacteria undergo cell division, not only FtsZ but also other proteins bind to FtsZ and work cooperatively. Although these proteins are densely packed, they are constantly in motion, and the question of how they can move without colliding into each other has long been a major mystery.
Therefore, to address this, we used cryo-electron microscopy to analyze the structure of the complex formed by ZapA and FtsZ, and we were fortunate to observe it at an exceptionally high resolution of approximately 2.7 Å. As a result, we found that ZapA binds to multiple FtsZ molecules, forming an overall structure resembling a “ladder” (Fig. 4).
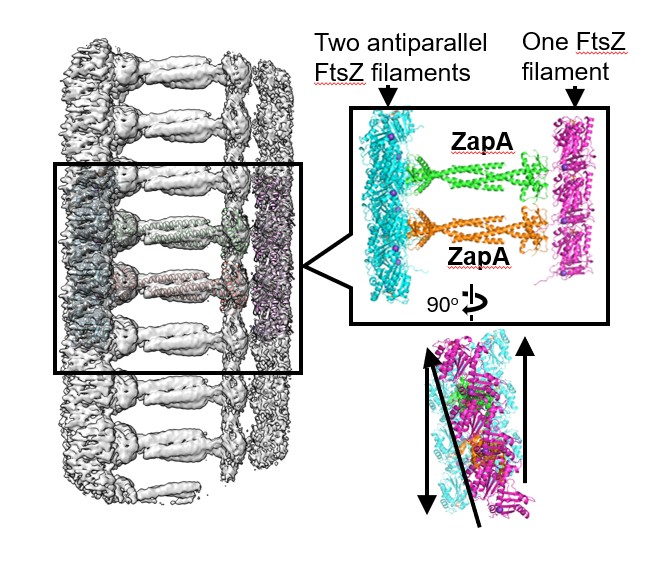
Fig. 4. A state where ZapA bundles FtsZ filaments.
What was particularly intriguing was that repulsive forces arise between the negatively charged regions of the bundled FtsZ filaments. Thanks to this repulsive force, we were able to demonstrate that even in a densely packed state, the proteins can move freely without interfering with one another.
Next, we initiated a joint research project with Professor Takayuki Uchihashi of Nagoya University and used high-speed atomic force microscopy (AFM) to observe, in real time, how ZapA binds to FtsZ. We found that ZapA binding is not static; rather, it is highly dynamic, repeatedly associating with and dissociating from FtsZ over short timescales.
In other words, during bacterial cell division, proteins repeatedly assemble and reassemble, much like scaffolding at a construction site, while the system as a whole continues to function effectively.
International reception
To be published in a top-tier journal like Nature Communications means that, as a researcher, you have secured a foothold in the highest echelons of research. I was certainly delighted when our papers were accepted.
Having said that, this journey was by no means easy. In fact, the two papers mentioned above each took nearly a year to be published. This is because multiple reviewers (i.e., researchers who evaluate papers blindly) provide various comments on the paper’s content, and each one requires an appropriate response. If even one reviewer feels the response is insufficient, publication is often withdrawn. We proceeded very cautiously—like walking on thin ice—repeating additional experiments again and again, and only then were we finally able to convince all of the reviewers.
I feel that the response has been excellent not only in Japan, but internationally as well. When our paper was published in 2023, I received an email saying ‘Congratulations!’ from a renowned professor, but at the time I still felt we had a long way to go.
Then, when we successfully published a second paper in the same journal in 2025, I received many congratulatory messages, and even offers for joint research from prominent researchers overseas. That really made me realize how important it is to publish your work in peer-reviewed journals. It feels like I’ve finally managed to stand, if only just barely, on the edge of the world’s top level.
Molecular interface control: A new approach to non-active sites
What these two papers have in common is an approach called “molecular interface control.”
Previously, it has been standard practice to directly modify the active sites of proteins, which are the central regions where chemical reactions occur.
However, we have continued our research with the perception that it may be possible to alter the overall function of a protein by modifying the molecular interfaces located away from the active sites, or by attaching artificially engineered molecules.
The inspiration for this molecular interface control approach came from a joint research project with Professor Hiroshi Fukayama of Kobe University on RuBisCO, a CO₂-fixing enzyme (Fig. 5). RuBisCO is the central enzyme responsible for fixing CO₂ into organic compounds in the Calvin cycle of photosynthesis. It is a large enzyme composed of eight copies each of the core component RbcL and the helper-like RbcS, which supports and regulates it.
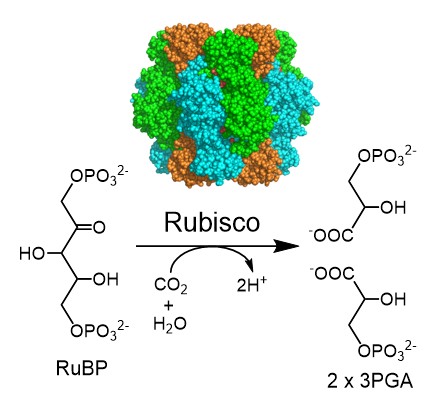
Fig. 5: Structure of RuBisCO (top) and the reaction it catalyzes. Rubisco is composed of eight RbcL units (shown in green and light blue) and eight RbcS units (shown in orange).
Enzymes possess active sites that catalyze chemical reactions; however, despite RuBisCO’s active site being located on the RbcL chain, simply replacing the RbcS chain somehow increased the activity of the RuBisCO enzyme. (Mol. Plant, 13, 1570 (2020)). This was a surprising finding.
It may be something like pressing a pressure point on your foot when you have a headache. Previously, researchers have directly stimulated the ‘pain point,’ that is, the active site, but we realized that by stimulating a distant molecular interface—like a pressure point on the foot—it might be possible to improve the function of the entire enzyme.
In addition, in collaboration with Professor Yoshihisa Nakazawa of Tokushima University and Dr. Hiroyuki Kajiura of the University of Osaka, we found that an enzyme called TPT, which is capable of producing biopolymers, is structurally very similar to another enzyme, FPS, which cannot produce polymers. Yet simply altering regions far from the active site enabled polymer production (Fig. 6; Commun. Biol., 4, 215 (2021)). This, too, was a surprising finding.
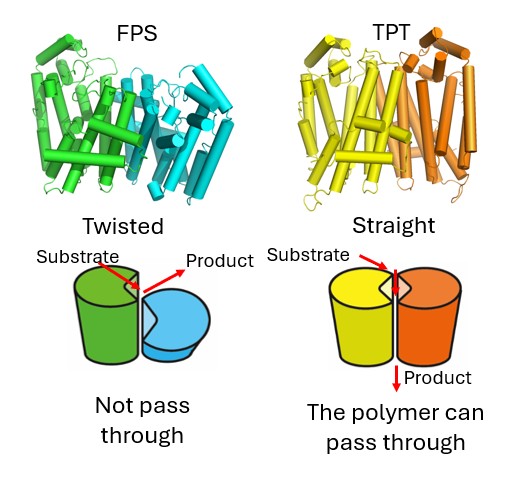
Fig. 6: Structures of FPS, which cannot produce polymers (left), and TPT, which can produce polymers (right). TPT has distinctive amino acid residues at its molecular interfaces, which relieve molecular twisting and create a tunnel through which the polymer can be released.
Since we were able to publish these studies consecutively in 2020 and 2021, I felt there was something particularly interesting there. Both studies showed that changing molecular interfaces led to changes in function, and that’s what prompted me to explore various possibilities under the key concept of molecular interface control.
Developing new treatments to address the problem of drug-resistant bacteria
Since my time in graduate school at the University of Osaka, I have been studying Co₂-fixing enzymes. Around 2010, however, I began to feel that I wanted to develop my own distinctive research topic, so I decided to focus on FtsZ—a bacterial cell division protein that is also a target for the antimicrobial drug development mentioned earlier—and I have pursued this line of research ever since.
Bacteria are all around us, including those in the gut and on the skin. Although they are invisible to the naked eye, bacteria are the most abundant organisms, with more than 100,000 species said to exist on Earth. When we become ill, harmful bacteria such as Staphylococcus or Klebsiella pneumoniae actively divide. This means that if we can design compounds that stop bacterial cell division, they can become effective medicines to cure diseases. For this reason, I wanted to investigate the mechanisms of bacterial cell division in detail.
At present, drug-resistant bacteria such as MRSA (methicillin-resistant Staphylococcus aureus), against which conventional antibiotics are ineffective, have become a global concern.
We believe that by understanding the fundamental mechanisms of bacterial cell division, our research has the potential to lead to the development of entirely new therapeutic approaches based on novel principles. In fact, in collaboration with Professor Daniel S. Pilch of Rutgers University in the United States, we successfully developed compounds in 2017 and 2023 that specifically bind to FtsZ and inhibit cell division in MRSA (Fig. 7; ACS Chem. Biol., 12 (7), 1947–1955 (2017); ACS Chem. Biol., 18, 629–642 (2023)). Because these compounds operate based on mechanisms different from those of conventional antibiotics, they are less likely to induce the emergence of new bacterial resistance.
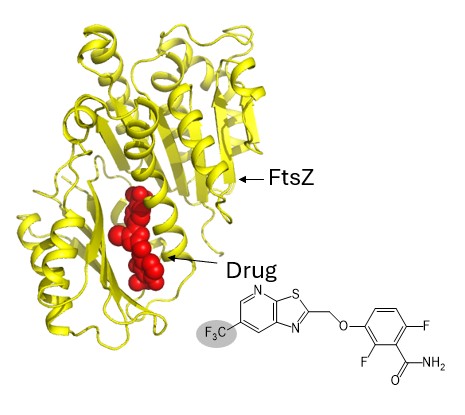
Fig. 7: A drug binding to FtsZ. Structural analysis will enable the development of drugs that bind more strongly, i.e., more effective drugs.
Further analysis is currently underway, and we hope to contribute to the development of new medical technologies with fewer side effects.
A discovery in plant photosynthesis: Potential implications for climate change mitigation
Another area that I’m particularly excited about right now is our research on RuBisCO.
The concentration of CO₂ in the atmosphere has now reached approximately 420 ppm (as of 2025), and this is considered a major driver of climate change.
Plants absorb CO₂ and produce oxygen through photosynthesis. Given the current rise in atmospheric CO₂ levels, plants should, in theory, have access to more nutrients and therefore grow more vigorously. However, for reasons that are not yet fully understood, the growth of many plants has plateaued. If we can somehow get rid of that limiting factor and allow them to consume more and more CO₂, they should be able to grow more rapidly and convert more CO₂ into oxygen. Results are beginning to emerge that could contribute to climate change mitigation in exactly this way.
Although this work is already publicly available as a preprint prior to journal submission (bioRxiv, 631008 (2025)), we found that modifying a molecular interface located away from RuBisCO’s active site led to a modest increase in RuBisCO activity, and as a result, the model plant Arabidopsis thaliana grew to about 1.3 times its ordinary size. This was something I had thought would be impossible to achieve during my own research career, and it has been the most surprising result so far.
We learned for the first time that even a very small improvement in enzyme performance—an increase of about 1.1-fold in catalytic rate—can have a major impact on plant growth. If we are able to further improve RuBisCO’s catalytic rate to 1.2- or 1.3-fold, the impact could be even greater. There may also be potential to apply this approach to other plant species as well.
We were told this was impossible, but luckily for us, we got the results, so I am glad that I have persevered in this research into RuBisCO.
The significance of insisting on doing things differently
I believe it’s important to do things differently from others and not follow trends.
In fact, we received the Crystallographic Society of Japan’s Academic Award in 2022 for our work on RuBisCO, so I think it has been academically recognized. In addition to RuBisCO, I have steadily pursued research on another CO₂-fixing enzyme called PEPC. However, practically speaking, I think it was considered “useless.”
For example, another study previously published in Nature showed how the authors were able to improve RuBisCO, but the plants expressing the enzyme ended up becoming very small, so many researchers felt that it was pointless to study RuBisCO. Nature is an extremely prestigious journal where researchers might be fortunate to publish just one paper in their academic career. Given that results like these had already been reported by others, I was honestly quite pessimistic about being able to produce meaningful results myself.
While I always had the belief that if we could improve RuBisCO and actually improve plants, the research would finally gain recognition, it was also true that funding for RuBisCO research was extremely limited. I was close to giving up, but I continued to pursue my research on a very small scale.
Then my findings completely overturned the previously held belief, which genuinely surprised me. Professor Shinichi Arimura and Dr. Issei Nakazato of the University of Tokyo had developed a new genome-editing technology, and together with Associate Professor Wataru Yamori—also of the University of Tokyo—they began discussing the idea of trying to modify RuBisCO using this technology. When they wondered who else was studying RuBisCO, they happened to find my work, so they reached out and invited me to collaborate with them. Where I am today is thanks to the success of this collaboration.
It was a research topic that went unnoticed for many years, but precisely because I continued with it, I was able to put it to use when the technology finally caught up. I’ve come to truly appreciate, through personal experience, the importance of doing things differently.
A sense of crisis about climate change: The driving force behind the research
The driving force behind my research is, above all, a strong sense of crisis about climate change. I love nature and often go hiking in the mountains on my days off, but the extreme heat of recent summers feels abnormal to me. I wonder what will happen to the Earth we live on.
It is predicted that the global population will reach 9.7 billion by 2050. On a planet where further warming is a serious concern, the greatest challenges over the next half-century will be how to sustainably supply sufficient food and energy to support humanity and how to contain the spread of infectious diseases exacerbated by climate change. My research aims to build a foundation for addressing these critical challenges in global environmental science.
Through molecular interface control, I hope to develop new enzymes, organisms, drugs, devices, and technologies, to expand collaborative research networks, and to actively communicate these outcomes outside the university.
Gratitude to Ritsumeikan and the serendipity born from collaboration with students
I came to Ritsumeikan University from the University of Osaka in 2015, and in 2022, I was selected as a RARA Associate Fellow. Since then, my research environment has changed significantly. I feel fortunate in many ways, and while I still have much to learn, I believe I am gradually producing results.
Serendipity often begins with chance events, mistakes, or things that were never intended. Through my collaborations with students at Ritsumeikan, I have experienced many moments of serendipity.
When I was at the University of Osaka, the students I worked with in my lab tended to carry out experiments precisely, following my instructions to the letter. As for myself, I am actually quite aware that my predictions often turn out to be wrong, and that many of the ideas I propose are not necessarily correct.
At Ritsumeikan, however, working with students who have diverse personalities—sometimes even ‘mischievous ones’—has led to unexpected discoveries. In fact, surprising results that emerged from miscommunication between us, or from students’ unforeseen actions, directly contributed to the success of the TPT research in 2021. That work ultimately led to my selection as a RARA Associate Fellow, which still feels somewhat surreal.
That’s why, when I work with students now, my stance is that it’s okay if they don’t follow my instructions to the letter, or even if they decide to try something completely different. Of course, there are cases where I do ask them to carry out specific experiments, but above all, I believe that the intuition of the person actually doing the hands-on work is the most important. That is my basic philosophy.
Aiming to cultivate a laboratory culture that does not fear failure
First of all, I ask students to set solid research topics so that they can steadily produce results and build confidence, but at the same time, I also encourage them to tackle more challenging topics. Each student usually works on multiple projects—sometimes juggling three or four at the same time. There are even cases where students take on ambitious topics and succeed across the board, and I am constantly amazed by their potential (Fig. 8).

Fig. 8: Members of my laboratory
I sometimes get the sense that students think, ‘There goes Prof. Matsumura again, saying something weird.’ But when a paper is published in a leading international journal and the students appear as co-authors, some of them later realize, ‘Oh, that was actually pretty impressive.’
Recently, many students have been choosing to advance to graduate school. All of the fourth-year undergraduate students I am supervising this year have decided to pursue further studies and to continue their research in my laboratory. Perhaps my passion for research aligns well with the motivation of my students.
Expanding international collaboration: A new student exchange program with Malaysia
Through my activities at RARA, opportunities for international research collaboration are expanding. In particular, a student exchange program with Universiti Putra Malaysia, where a student I supervised 20 years ago is now a professor, has become a reality thanks to the cooperation of the faculty and staff of the College of Life Sciences, and approximately 20 students per year participate in this program.
In 2019, I spent two weeks in Malaysia, where we organized workshops and a symposium on protein structure analysis (Fig. 9).
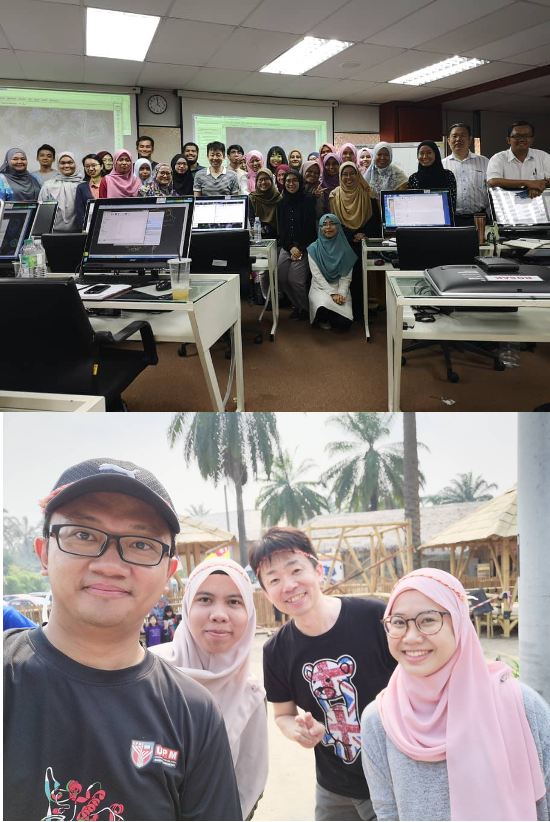

Fig. 9: Workshop at Universiti Putra Malaysia (top) and an excursion with program participants (bottom)
The widespread adoption of online meetings during the COVID-19 pandemic has significantly lowered the barriers to international collaboration, making it far easier to work closely with researchers around the world.
My message for early-career researchers
To the early-career researchers reading this article, I know it is not easy, but I encourage you to keep writing papers. As part of that process, aim to produce research that truly stands out—and, if possible, challenge yourself to submit your work to top-tier journals. Publishing even a single paper in a leading journal will allow you to see the world differently; publishing two will open the door to the world even more. One of the strengths of Ritsumeikan is that it allows researchers to attempt high-risk, basic research.
As researchers, however, we must pursue ideas that no one else is trying, and that can be a lonesome path. Even so, we must continue to challenge ourselves. Maintaining motivation is often difficult, especially when research does not produce results and it becomes tempting to blame the environment.
In such moments, I think it is important to have an attitude of proactively communicating with others so that you can be inspired by them, to be aware of your own ignorance and go to ask them about things you don’t know, and to support the growth of those around you so that you can consult with them when you need to.
Research that has been pursued over many years can sometimes flourish thanks to advances in technology. By persevering and not giving up, unexpected breakthroughs can emerge. Perseverance is key, so let’s keep working hard together.
Activities at RARA and my vision for the future
Being appointed as a RARA Associate Fellow has expanded my opportunities to collaborate with researchers from other fields, enabling me to move forward with truly interdisciplinary research. At a time when global population growth and the impacts of climate change are raising serious concerns, our mission is to help build the foundations for addressing key challenges in global environmental sciences in 50 years’ time, such as the sustainable supply of food and energy to support humanity and measures to combat infectious diseases.
With molecular interface control, I aim to develop new enzymes, organisms, drugs, devices, and technologies and to contribute to fields that are directly connected to the everyday lives of the general public. With gratitude for the environment I find myself in, I will continue to pursue ambitious research.
Acknowledgments
The research introduced here has been carried out with the cooperation of Assistant Professor Ryo Uehara, Ms. Chie Okamoto, and numerous students from the Structural Bioscience Lab, Department of Biotechnology, College of Life Sciences, Ritsumeikan University. I would like to take this opportunity to express my sincere gratitude to everyone involved.



